GROMACS is one of the most widely-used codes in all of HPC, providing key mechanistic and energetic insight into numerous biological processes. It provided one of the two major computational engines behind the distributed-computing Folding@Home project (https://foldingathome.org/). During the height of the global response to the COVID-19 pandemic, everyday computer users combined to provide more than one exaflop of computational power for understanding of the virus and investigating potential strategies (https://en.wikipedia.org/wiki/Folding@home#cite_note-8). GROMACS was also deployed for exploring potential COVID-19 drug candidates on Summit (https://www.olcf.ornl.gov/olcf-resources/compute-systems/summit/) the then-fastest-rated supercomputer in the world (see https://www.olcf.ornl.gov/2020/03/05/ornl-team-enlists-worlds-fastest-supercomputer-to-combat-the-coronavirus/ for details).
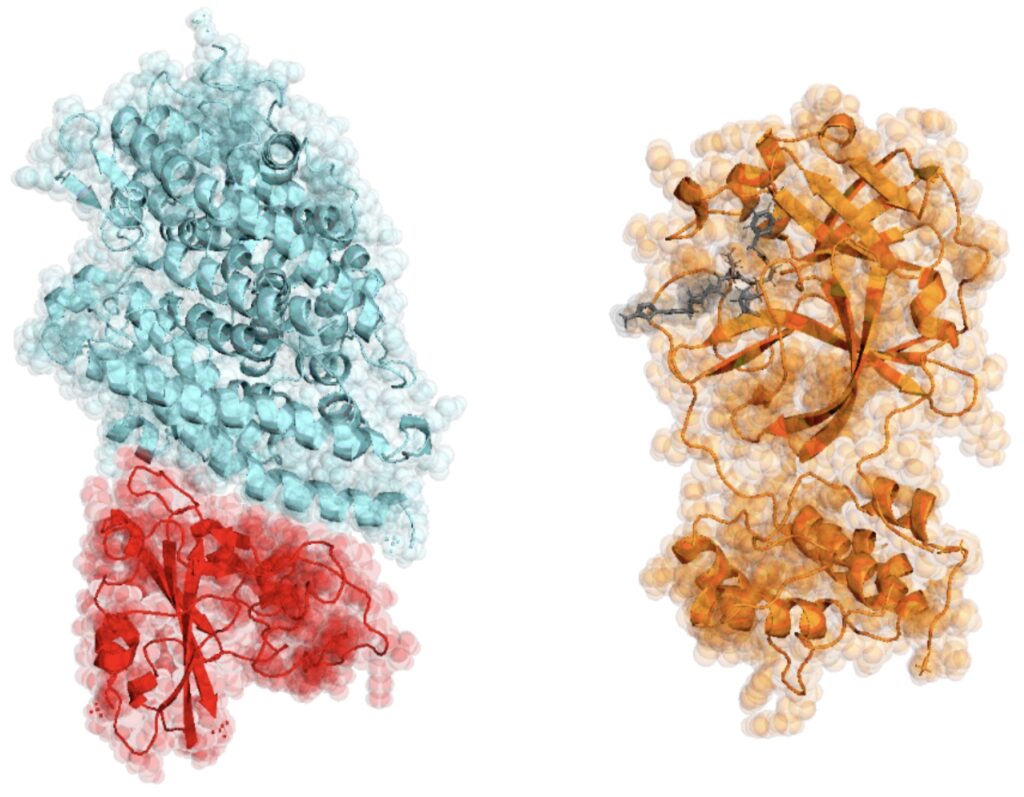
GROMACS is already ported to and runs fast on nearly every known high-performance computing platform, which is a tremendous achievement of software engineering (https://arxiv.org/abs/2006.09167). To routinely reach the exascale for many more kinds of biomolecular simulation problems, GROMACS requires continuing investment into four key aspects:
- increasing performance in computing a fixed-size trajectory on the same hardware,
- widening performance portability to new generations of accelerators and processors
- improving scalability of fixed-size trajectories to more hardware, and
- loosely coupling numerous trajectories with API-driven approaches such as swarms and ensembles.
ENCCS will contribute to all of these, as well as to the ongoing organizational and computational challenge of testing and integrating the necessary changes from the global development team. A key part of this approach is peer-based code review and requiring automated CI testing before accepting changes. These processes have proven their worth over time, and we plan to bring their benefits to the other codes under development at ENCCS.
For more information on GROMACS visit http://www.gromacs.org.
RECENT NEWS
[post_grid id=’651′]