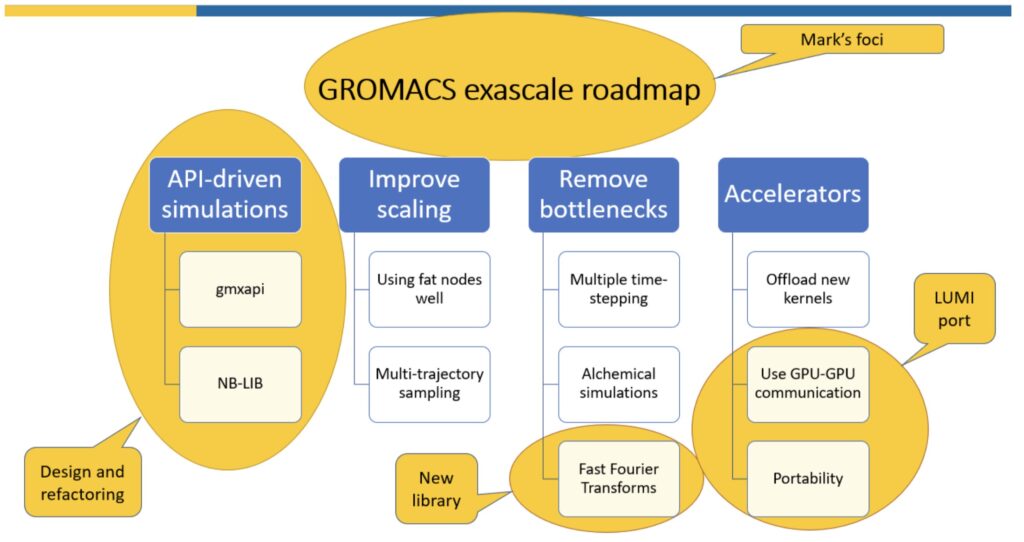
On Nov 6, the first ENCCS developer progress meeting was held. We took an in-depth look at the work Mark Abraham will do in Life Science, particularly with GROMACS. GROMACS is a mature widely-used code with a clearly articulated exascale roadmap (see https://zenodo.org/record/3563747). Mark’s work will focus on areas of that roadmap most relevant to Swedish researchers, particularly for using new HPC infrastructure such as the LUMI supercomputer (https://www.lumi-supercomputer.eu/).
Mark will
- Collaborate with AMD to find a suitable way to support the GPUs on LUMI, either via AMD’s vendor-owned HIP language or the upcoming standard SYCL language via hipSYCL (https://github.com/illuhad/hipSYCL),
- Contribute design and refactoring work to enable researchers to easily implement robust and performant API-driven workflows in Python (http://manual.gromacs.org/current/gmxapi/) or C++ (upcoming in GROMACS 2021)
- Develop a new modern C++ FFT library to provide an independent showcase of ENCCS development capabilities, while delivering performance enhancements to GROMACS, which uses small 3D FFTs not typically optimized well by existing libraries
For more information visit http://www.gromacs.org.
RECENT NEWS
[post_grid id=’651′]